 Interleaved files occur when the R1 and R2 reads are combined in one file, so that for each read pair, the R1 read in the file comes immediately before the R2 read, followed by the R1 read for the next read pair, and so on. This will extend the box to show three additional parameters that can be selected. If one wishes to change these parameters, click on the down arrow following Advanced in the Paired read library box. It will also infer the platform from the type of reads that it sees. The assembly protocol in BV-BRC assumes that the paired end reads are not interleaved and that the library creation was standard. This will move your file into the Selected libraries box. To finish the upload, click on the icon of an arrow within a circle. Repeat to upload the second pair of reads. Do not submit the job until the upload is 100% complete. Pay attention to the upload monitor in the lower right corner of the BV-BRC page. This will auto-fill the name of the document into the text box. Once selected, it will autofill the name of the file. 
Select the file where you stored the fastq file on your computer and click “Open”. This will open a window that allows you to choose files that are stored on your computer. Click on the “Select File” in the blue bar. This opens a new window where the file you want to upload can be selected. Click on the icon with the arrow pointing up. This opens up a window where the files for upload can be selected. To initiate the upload, first click on the folder icon.Īt the top of any BV-BRC page, find the Services tab and click on it The reads must be located in the workspace. To upload a fastq file that contains paired reads, locate the box called “Paired read library.” If PacBio and Illumina reads are available, both would be combined to generate the best assembly. Note that reads from different sequencing platforms of the same organism can be submitted in one job. What follows is a tutorial showing how to submit reads of various types for assembly and selecting parameters for the assembly algorithm. These differences necessitate different algorithms for assembly from short and long read technologies. Third generation technologies like PacBio and fourth generation technologies like Oxford Nanopore (called “long-read” technologies) provide read lengths in the thousands or tens of thousands but have much higher error rates of around 10-20%, with errors being chiefly insertions and deletions. Second generation sequencing technologies like Illumina (called “short-read” technologies) produce short reads on the order of 50-200 base pairs and have low error rates of around 0.5-2%, with the errors chiefly being substitution errors. 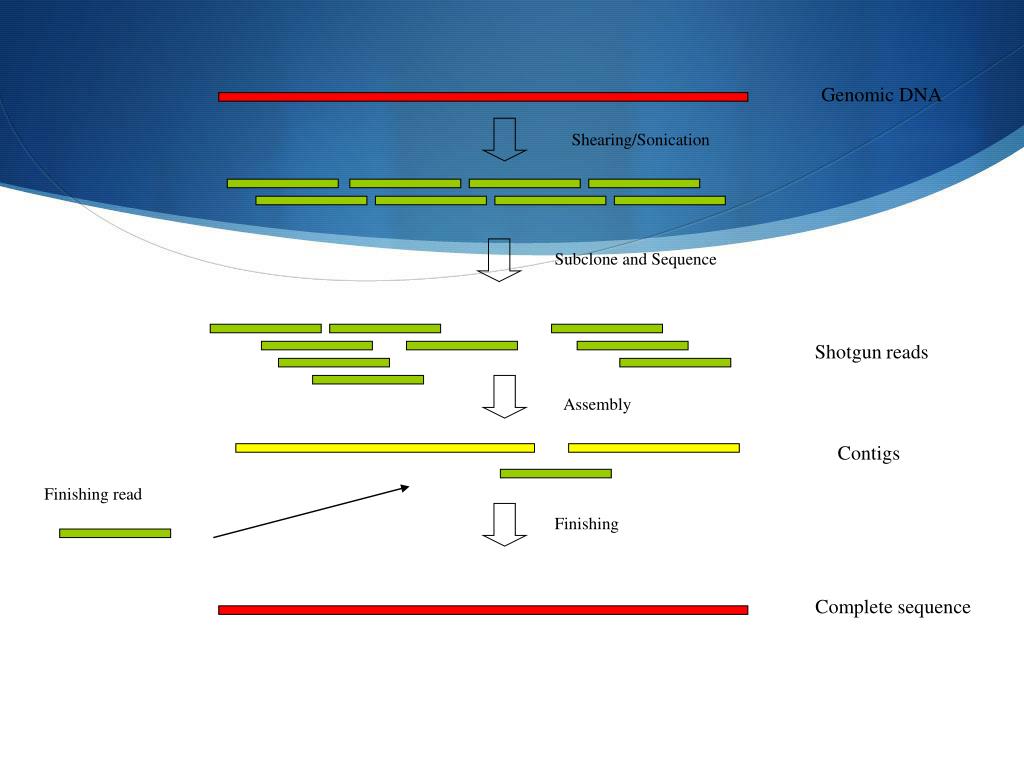 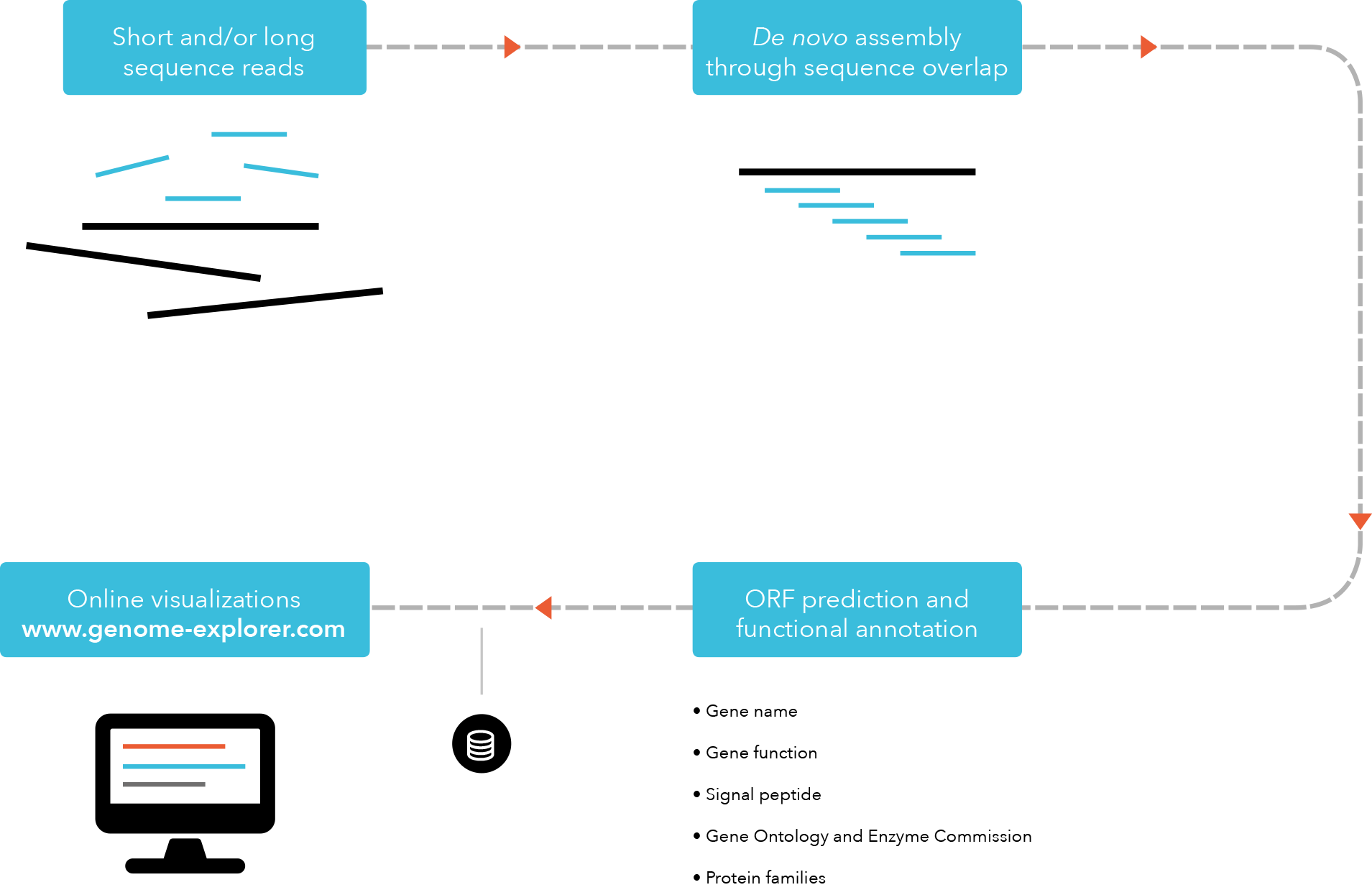
These are commonly used in bioinformatic studies to assemble genomes or transcriptomes.ĭifferent assemblers are designed for different type of read technologies. Typically, the short fragments, called reads, result from shotgun (random) sequencing of genomic DNA.ĭe novo sequence assemblers are a type of program that assembles short nucleotide sequences into longer ones without the use of a reference genome. This is currently needed as DNA sequencing technology cannot read whole genomes in one go, but rather can read small pieces of between 20 and 30,000 bases, depending on the technology used. Metadata-driven Comparative Analysis Tool (Meta-CATS)Ī genome assembly is the sequence produced after chromosomes from the organism have been fragmented, those fragments have been sequenced, and the resulting sequences have been put back together.SARS-CoV-2 Genome Assembly and Annotation Service.Comprehensive Genome Analysis (CGA) Service.Submitting reads that are present at the Sequence Read Archive (SRA). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
